Role of Ribosomal Protein bS1 in Orthogonal mRNA Start Codon Selection (Biochemistry, 2025)
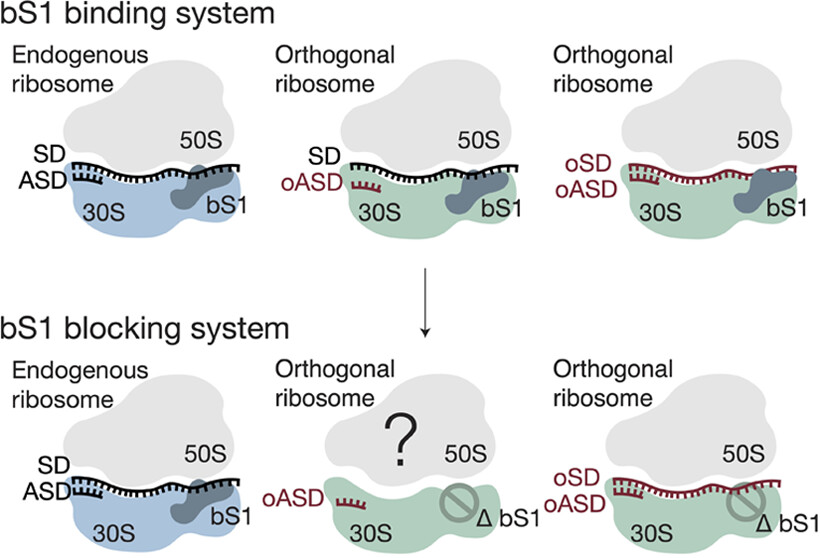
C-GEM researchers investigated whether preventing the binding of ribosomal protein bS1 could improve ribosome orthogonality. A current, common strategy for generating orthogonal ribosomes involves controlling mRNA and ribosome interactions by modifying the Shine-Dalgarno and anti-Shine-Dalgarno sequences, the sequence that sits upstream of the start codon… Read More »Role of Ribosomal Protein bS1 in Orthogonal mRNA Start Codon Selection (Biochemistry, 2025)