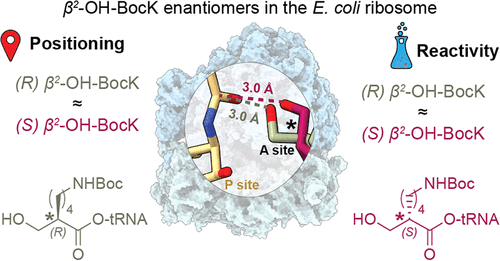
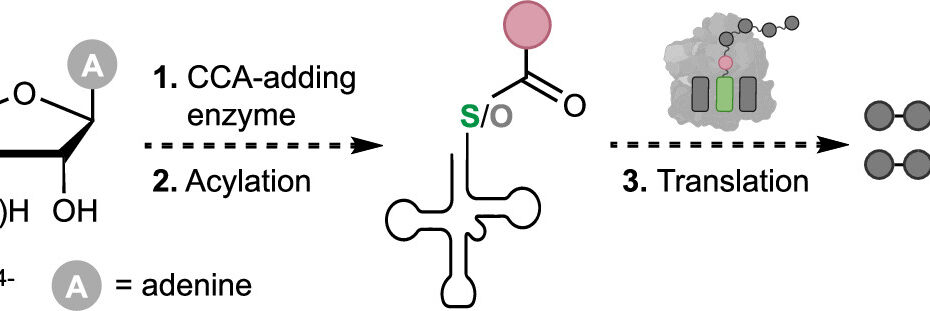
Co-Translational Incorporation of (R)- and (S)-β2-Hydroxy Acids In Vitro: A Structural and Biochemical Study on the E. coli Ribosome (J Am Chem Soc, 2026)
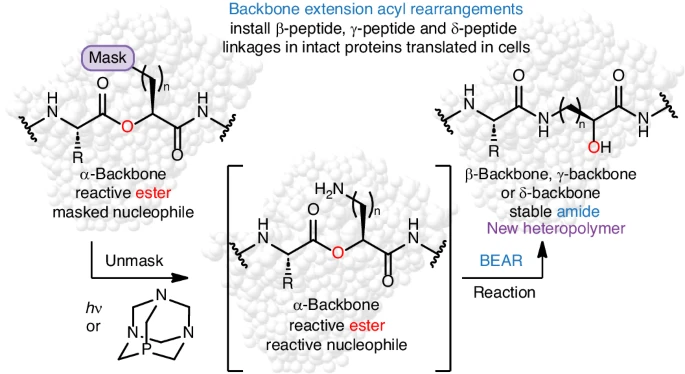
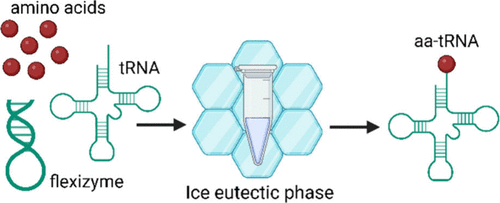
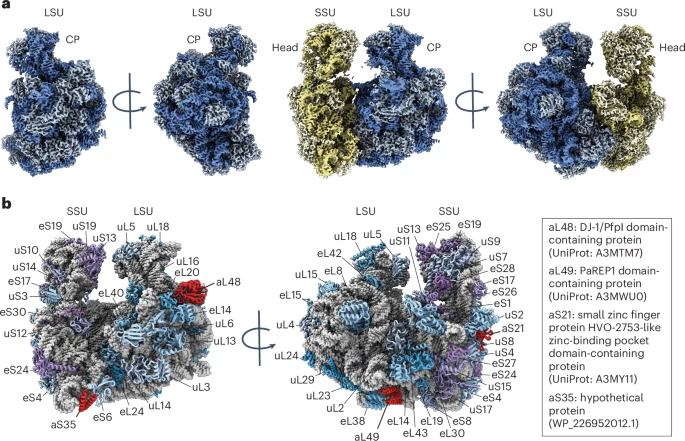
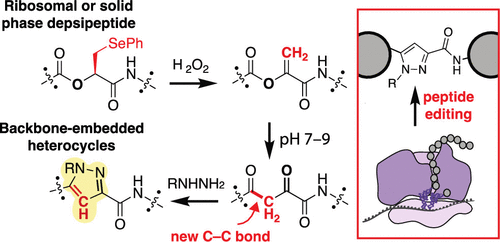
By Alexandra Kent The goal of C-GEM is to generate ribosomally synthesized, sequence-defined polymers using a diverse set of non-natural monomers. Although ribosomes are known to incorporate some unnatural monomers, the basis for observed differences in incorporation efficiencies of different stereoisomers remains poorly understood. In… Read More »Co-Translational Incorporation of (R)- and (S)-β2-Hydroxy Acids In Vitro: A Structural and Biochemical Study on the E. coli Ribosome (J Am Chem Soc, 2026)